HistoAtlas
HistoAtlas is a comprehensive resource mapping the morphological landscape of cancer across 21 TCGA cancer types. By extracting quantitative histological features from over 6,000 diagnostic whole-slide images, HistoAtlas reveals how tissue architecture relates to patient...
Cost / License
- Free
- Open Source
Platforms
- Online
Features
HistoAtlas News & Activities
Recent activities
- pierreantoine-bannier added HistoAtlas
 pierreantoine-bannier added HistoAtlas as alternative to Serial Cloner, CLC Genomics Workbench, SnapGene Viewer and Genophore
pierreantoine-bannier added HistoAtlas as alternative to Serial Cloner, CLC Genomics Workbench, SnapGene Viewer and Genophore
HistoAtlas information
What is HistoAtlas?
HistoAtlas is a comprehensive resource mapping the morphological landscape of cancer across 21 TCGA cancer types. By extracting quantitative histological features from over 6,000 diagnostic whole-slide images, HistoAtlas reveals how tissue architecture relates to patient survival, molecular subtypes, mutational profiles, and immune microenvironment composition.
Data Scope
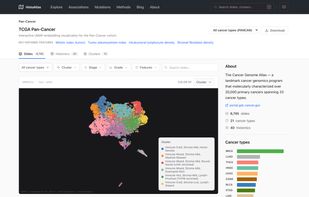
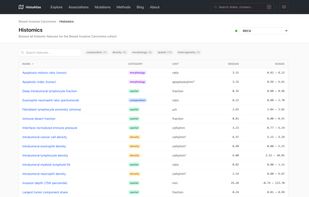
21 cancer types from The Cancer Genome Atlas (TCGA) Pan-Cancer Atlas 6,000+ H&E whole-slide images analyzed with deep learning-based cell and tissue segmentation 40 morphological features capturing cellular composition, tissue architecture, and spatial organization Multi-omic integration with mutations, copy number alterations, gene expression, pathway scores, and immune cell estimates
Key Capabilities
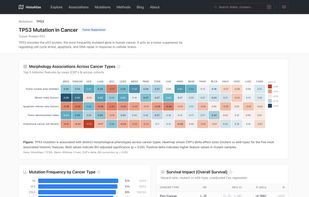
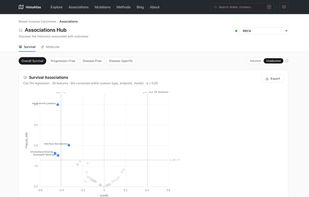
Survival analysis: Cox proportional hazards models linking morphological features to overall survival, with Kaplan-Meier curves and restricted mean survival time (RMST) estimates Molecular correlations: Spearman correlations between histological features and gene expression, pathway activity (GSVA), copy number alterations, and immune cell infiltration scores Categorical associations: Associations between morphological features and somatic mutations, molecular subtypes, and treatment response, quantified with Cliff's delta effect sizes Morphological clustering: Pan-cancer and cancer-specific clustering of slides based on morphological profiles, revealing shared histological phenotypes across cancer types Interactive exploration: UMAP embeddings, slide-level deep dives, and cluster characterization through an interactive web interface